Guest blog by Dr. Alex M. Clark, Molecular Materials Informatics, Inc.
What do you get when you combine a company with an online collaborative chemistry sharing platform, a startup dedicated to producing cutting edge apps for chemistry, a consultant with deep domain knowledge and lots of ideas, a researcher with access to valuable data, and a generous grant for helping to find cures for neglected diseases?
Many things, but the first one is a mobile app called TB Mobile, which is available to anyone for free on the iTunes AppStore and for Android on Google Play.
The app is one of the deliverables from a grant funded by the NIH which involved using existing information about small drug-like molecules that are known to bind to one or more of the proteins of Mycobacterium tuberculosis. This information, collated for small molecules and targets by CDD and on pathways and bioinformatics data by Dr. Malabika Sarker at SRI International, was assembled using the online data management tools provided by Collaborative Drug Discovery. The information in the app includes more than 700 chemical structures, each of which has associated protein targets, pathways, essentiality and human ortholog information, with a lot of links to literature references.
But this information alone is not necessarily easy to access by everyone involved in the global hunt for new and better cures for this resurgent disease. Anticipating this, Dr. Sean Ekins, CDD VP of Science, included a section in the grant proposal for appifying the results, in order to magnify the audience for this valuable resource, and make it as easy as possible for anyone to make use of it. Once the data was complete, CEO Dr. Barry Bunin commissioned my company, Molecular Materials Informatics, to take the raw data and bundle it in the form of a simple, intuitive and visually pleasing app, and make it available to anyone who has an iPhone, iPad, iPod or an Android-based mobile device.
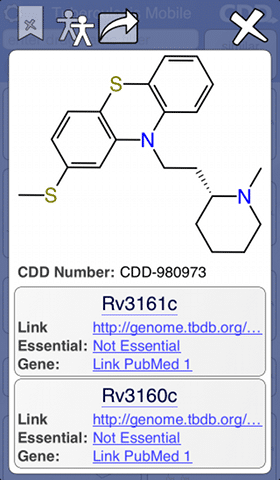
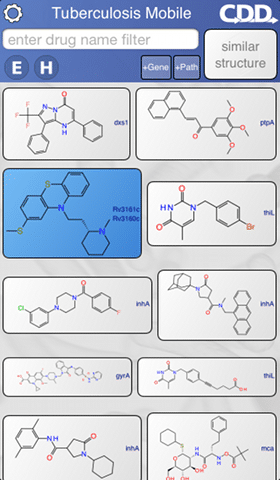
As anyone with a mobile device knows, installing a free app could hardly be easier: click on the link, press install, wait for the download to finish, then tap on the icon to launch it. The TB Mobile app is about as simple as it gets: the front page displays a list of structures, and swiping up or down allows the list to be browsed. Tapping on any of the structures brings up a detail page, which reveals all the known information about the compound, as well as a number of links that can be followed for more detailed reference articles.
The control bar along the top provides ways to organize the data, by searching for name, limiting to specific disease targets, or ranking by similarity to a structure which can be sketched within the app. In short, this simple to use app places a lot of very useful data about potential tuberculosis drug candidates and other compounds that have been tested in vitro in the hands of anyone who has a mobile device running either of the most popular platforms. One workflow might be for a hit from a phenotypic screen to be used as an input in order to suggest potential targets. This has been termed target fishing and the features in the app enable this, albeit as a rudimentary first attempt before applying more powerful tools or experimental validation.
Because the data for the app originates from a CDD vault, users of the online collaborative service have access to the data in full, and can integrate it into any of their workflows within the CDD Vault. The appification represents an effortless entry point for casual users, while the full service provided by CDD is a natural progression for more involved workflows. The TB Mobile app is the first of these adventures, and it only has to help one scientist get closer to finding a cure for tuberculosis to justify its value many times over. But it also represents a proof of concept experiment for making open data available to the widest possible audience. We believe that apps are a great way to do this, and by all means help prove us correct by downloading and using it!
This blog is authored by members of the CDD Vault community. CDD Vault is a hosted drug discovery informatics platform that securely manages both private and external biological and chemical data. It provides core functionality including chemical registration, structure activity relationship, chemical inventory, and electronic lab notebook capabilities!
CDD Vault: Drug Discovery Informatics your whole project team will embrace!


