Update structureless molecules en masse (For vaults that use CDD's molecular registration system)
If you work in a vault with a registration system, you previously had to manually update each structureless molecule when the structure became available. That’s fine for the one-off compound, but a drag for a whole compound library. Now you can bulk update structures just by importing a file! We’ve even built in batch salt and formula weight updating, if you wish to update those, and all you have to do, is include a batch identifier in your import file mapping. The best part is that any user with sufficient privileges to edit existing molecules can do this!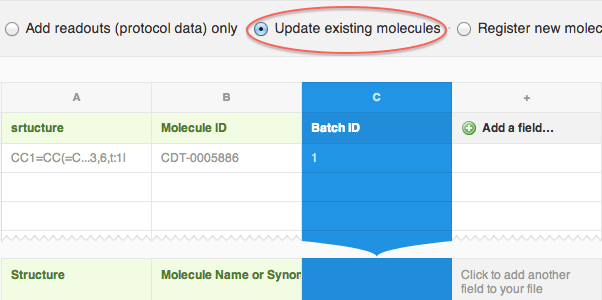
- Create an import file with the necessary details:
- To add structures, but not update existing batches, include:
- Structure (SMILES, MOL)
- Molecule ID
- To add structures and update existing batch names and salts, include:
- Structure (SMILES, MOL)
- Unique Batch ID
- Molecule ID (optional if unique batch ID is included)
- Batch Name (optional if unique batch ID is included)
- To add structures, but not update existing batches, include:
- Import the file to CDD and choose to “Update existing molecules”
- Map structure and the molecule and batch IDs just like you would when registering new molecules, and proceed with the import
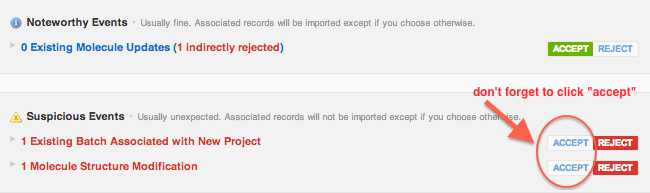
Batch name not required for molecule updates on file import (For vaults that use CDD's molecular registration system)
When updating molecule fields, such as user-defined fields or synonyms, there's no reason a batch name should be required. So when making updates to molecules in vaults with chemical registration systems via file import, a batch name doesn't have to be mapped on step 2.Simpler ownership of data imports
The user who processes a file will be the effective owner of the data, regardless of who uploads the initial file or who clicks commit. This same user will be the data owner listed on all imported data.Read-export user role update
We’ve modified the read-export user role to include both the ability to export search results and to download attached files. Read the full user-role descriptions.Additional options in “Default display” settings
Display Defaults (introduced about a year ago) have been extended to include global results settings: Detail level, Displayed readouts, and dose-response plots scale. You can read all the details, but briefly:Detail level determines how many rows of data are displayed per each molecule, with the “details” level being the most extended view, and “summary” view showing just one row of data. Displayed readouts controls what sub-set of protocol results is displayed for each matching molecule. Dose-Response plots scale controls the y-axis scale, which can be adjusted to a standarized scale across all plots in a single run, or individually for each plot.
Vault administrators may change the defaults on the Settings-> Display Defaults tab, which will apply to all searches performed in the vault. Individual search settings may still be adjusted by any user under “Customize your report”.Other posts you might be interested in
View All Posts
CDD Blog
2 min
April 22, 2024
Recorded Presentations: CDD 20th Anniversary User Group Meeting
Read More
CDD Blog
9 min
April 19, 2024
Drug Discovery Industry Roundup with Barry Bunin — April 19, 2024
Read More
CDD Blog
2 min
April 19, 2024
CDD Appoints Yasushi Hamagashira as Head of Sales and Marketing for Japan
Read More


