Readout definitions: more flexible averaging
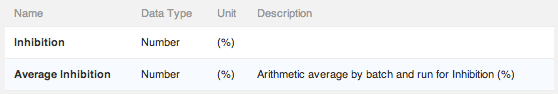
You can now choose a different level of aggregation for each average readout, whereas previously the aggregation level was uniformly set for all readout definitions in the vault.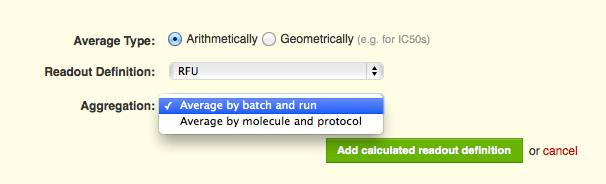
- Batch/Run is good for replicate data in a single experiment (run). Replicated data from the same batch of a molecule will be averaged together for each run of the protocol.
- Molecule/Protocol works well for generating summary results for molecules tested in a protocol. Data from all batches of a molecule will be averaged together across all runs of a protocol.

Percent of negative control normalization for dose-response calculations
While positive controls are almost always a good idea, some assays are run without them. Until now, dose-response calculations required both a positive and a negative control if the raw data was to be automatically normalized within the CDD Vault. In this release you can normalize your dose-response data as percent of negative control. You will see the new option when you define a dose-response calculation in your protocol: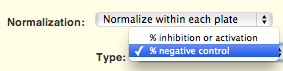
Molecule description has moved
The “description” field, which was previously part of the core molecule definition, is now just another user-defined field. If you had any data in the description field, it was automatically migrated.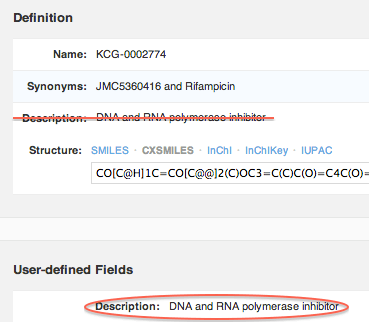
Other posts you might be interested in
View All Posts
CDD Vault Snack
4 min
April 15, 2024
Vault Snack #23 – New Interface for Creating Protocol Readout Definitions
Read More
CDD Blog
9 min
March 26, 2024
Drug Discovery Industry Roundup with Barry Bunin — March 26, 2024
Read More
CDD Blog
2 min
March 26, 2024
Recorded Webinar: Trends & Better Practices for the Use of Gen AI & LLMs
Read More


