March 8, 2010
Guest Blog: Dr. Jonathan Baell, Medicinal Chemistry Group Leader, CRC for Cancer Therapeutics at WEHI
We at CDD wish to bring attention to useful developments to the community. In this case it was a recently published J. Med. Chem paper brought to our attention by our SAB member Christopher Lipinski titled "New Substructure Filters for Removal of Pan Assay Interference Compounds (PAINS) from Screening Libraries and for Their Exclusion in Bioassays." We think this paper will help the drug discovery community avoid wasting valuable resources on red herrings. We're delighted to sponsor a guest blog entry from the author, highlighting some of the more personal considerations behind the scientific drivers courtesy of Dr. Jonathan Baell at the Walter and Eliza Hall Institute of Medical Research in Australia. In addition to highlighting this important work, we invite others from the community to contribute to this or other timely topics.
“Patent applications and issued patents are now as valuable for tenure decisions/standing as quality manuscripts in the academic world.”
— Jeff Aube 2009
If translational research is your caper, this quote would surely be music to your ears, because a little aspartame to reward the difficult journey of academic drug discovery is always welcome. In short, recognition of the difficulties of value-adding research appears to be gaining traction. High time, because you know that your compound is only valuable if protected by a patent and that this process can hinder timely publication - so a different metric to judge your work is sensible.
And by definition, a patent implies novelty and utility and, by inference for some, ingenuity as well, right? What better a way to tangibly show the tax payer where their money is going? This notion should be rewarded and encouraged, as should translational research.
There’s a problem, of course: any patent attorney with a skerrick of initiative could probably file a provisional patent on the analgesic or antipyretic properties of carboxyphen-2-yl acetate and happily take a few bucks in return (a few for the provisional, but later it turns ugly). Patents aren’t peer reviewed and unlike papers, aren’t required to be written in a way that allows ready assessment of the scientific ingenuity involved. And it’s not until perhaps many years later after a patent is granted that there is a reasonable guarantee the work is even novel and of utility. This could make for a messy metric in judging translational track records. A real stew. So the obvious solution is to value more those patents that are accompanied by a publication, because peer review is the most expedient process for assessment of real scientific advances, right?
Regrettably no, and right now the academic chemistry-biology nexus is in the cauldron. Like any robust marriage, this wonderful and lauded union is not without its problems and right now, these are going public.
Specifically, I’m talking here about the discovery of compounds that light up your primary competition assay, then confirm in your secondary binding assay, and finally readout successfully in your cell-based assay. A bit of a lit search reveals healthy evidence of biological activity in related compounds – always a good sign [as opposed to those classes where available analogues are numerous but devoid of biological literature. A definite yellow card. See this useful link: Christopher Lipinski, PhD (ex-Pfizer, Melior Discovery)]. And you can even see similarities to known drugs. So with a flourish, you make a few analogues into clear IP space, file a composition of matter patent, promptly publish in a reasonable journal and voila! – the translational track record is starting to look good.
But there’s a problem. A big, problem. There’s a good chance these compounds are actually worthless and simply outwitting you with false readouts in any assay you throw at them. Even if your screening set has been filtered free of known reactive groups, your next cure for cancer might nevertheless be protein-reactive and with little hope of development.
Take alkenylrhodanines, for example. They’re often coloured, can chelate metals and react with proteins in multiple ways – sidechain nucleophiles and peptide backbone – with more than one reactive site on the rhodanine itself! In some rhodanines but not in others, reactivity may be light induced. Worse still, they may only register as a hit against your protein and not another used as a selectivity control. Basically, these nuisances can confirm in multiple different assay protocols and appear convincingly as a real hit, ripe for progression. Being readily available, they are common in sets of in silico screening hits and will similarly present themselves as confirmed actives – and appear to simultaneously validate the in silico screen. Whether from wet or dry screening, these compounds can regrettably select themselves for progression over other weaker compounds that are more intrinsically optimizable that may (or may not) be present.
In the early days of HTS, the pharmaceutical industry no doubt would have made acquaintances with many of these screening hits and for many years have necessarily recognized these as “no-go” options. The academic researcher has not had this historic luxury and the whole situation is exacerbated by the relatively sudden facile access to HTS by academics.
I should know. These super smart false positives (SSFPs, anyone?), these pan assay interference compounds (PAINS perhaps? – hmmm, definitely more palatable) have caused us some grief not so many years ago. For that reason, we have taken some effort to identify these PAINS and my colleague Dr Georgina Holloway and I have recently published our findings in the Journal of Medicinal Chemistry, making all our results freely available. We are pleased but also humbled by the interest this work has received.
Rhodanines are just one of over 400 classes of potential PAINS that we identify. But most of these are singletons – rare chemistry – and the academic researcher should not be intimidated into learning to recognise the worst offenders; these are dominated by only a handful of well-defined and readily recognized classes.
So what do you do if you find yourself with a PAIN that is presenting promising biological data? My instinctive response is to enact the “reflex de in-laws” protocol (rather than the less effective accord): eschew with alacrity and regard close relatives with automatic suspicion.
But this response needs to be strongly qualified: currently, if it is one of the lesser known PAINS and while it is still lesser known, it might be useful to try to answer the question “if this may be interfering with my assay, how is it doing so?” In fact, I would welcome the academic drug discovery community to help us all understand PAINS better in order that we may better define their various current and future classes. I would like to know, for example, how the fused tetrahydroquinolines that we discuss may be protein-reactive. The first person to find this out could readily publish their results: we have a PhD student specifically investigating another putative PAIN to discover more about the SAR of the particular class under scrutiny.
For some of you reading this it may be too late and your student may be finishing up and ready to publish. I think everyone is sympathetic to such needs, but at the very least, perhaps an allusion to potential problems could be penned in your manuscript? And maybe not paint your PAINS as promising leads?
More generally, any publication reporting compounds as genuine assay positives should ideally be accompanied by a swathe of convincing data. This includes evidence of specific and reversible binding, clear SAR in say a dozen good probes and if possible a functional biomarker response (biochemical; cellular good but not necessary). If the literature is full of similar compounds as hits for different targets in different assays, the red flag goes up and the player is sent from the field. Ironically, an absence of such literature should not be taken as reassurance because you might simply be dealing with rare chemistry (see also the link above for other reasons). Something in between, where the literature is not necessarily screening-based but further downstream, is just about right.
Note that I am entirely relaxed about publishing screening hits that are structurally suspect (although I’ve had enough of the rhodanines and a few others) if this point is made, or at least if the point ISN’T made that they are highly optimizable. This to some extent is still a useful public awareness exercise and you will see future work from our group on such aspects.
In a practical sense, the easiest way to identify PAINS (for users of Sybyl) is to run dbslnfilter using the slns provided in our paper (or to contact me for the text files if conversion from the pdf proves troublesome) as detailed in the experimental section (many thanks to Craig Morton for picking up an error here which is now corrected in the current ASAP article). For users of SMARTS, we hope soon to be able to provide the filters in this form also thanks to the efforts of nascent colleagues Ian Watson, Lakshmi Akella, Rajarshi Guha and Wolf Ihlenfeldt.
The current aim is to make these filters fully functional at CDD.
In our paper, we also provide structural representations of all our likely and putative PAINS in an extensive listing in the supplementary information (Figure S1). While less accurate, this is useful for those who can’t deal with the text filters. But their role is greater than simply this: they provide a key to recognising PAINS not recognised by the strict definitions of the text filters. This requires some explanation that was not discussed in our PAINS paper.
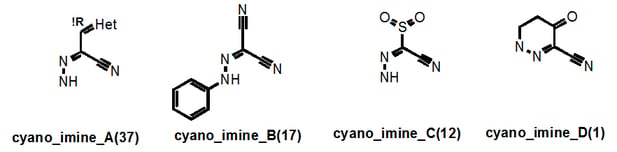
Let’s take a look at the “cyano_imine” family of PAINS. These are easy to find in supplementary Figure S1 where the PAINS are listed in alphabetical order:
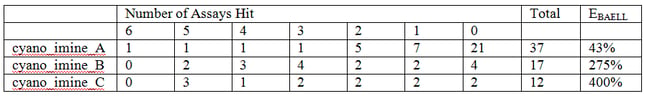
Here the numbers in parentheses tell us the number of these that are in our library. This number then usefully links in to supplementary Table S7, which filters in order of family size and so on page S24, the exact sln definition of cyano_imine_A is readily located. Supplementary Table S9 may be of more general use as this Table contains the breakdown of the "assay counts." This table also lists the PAINS in order of how frequently they occur in our library. So it’s easy to get the following data:
Here it can be seen by that cyano_imine_B for example is very nasty and the percentage that hit between 2 and 6 assays as opposed to none (the “EBAELL” value) is 275%!! Anything over 30% is bad – and cyano_imine_C is even worse. It’s not hard to speculate that attack on the nitrile or the imine by protein nucleophiles could be the problem.
So why did we not filter such groups out in the first place? Well for a start, cyano_imine_B is present in Levosimendan though admittedly the PK of this iv-only active drug is shot if oral activity and a long half-life had been required. (http://indianpediatrics.net/july2009/593.pdf). Secondly, it is very hard to envisage all combinations of unpublicized functional groups until you find them in your assays. Either way, we now do not want these as screening hits and do not want these in our library.
So how did I name these classes of PAINS? Well, once all classes had been structurally defined, it was clear that there were recurring themes: certain anilines (anil_), alkenes (ene_), nitriles (cyano_), imines (imine_), hydrazones (hzone_), pyrroles (pyrrole_), miscellaneous heterocycles (het_), sulfur-containing heterocycles (het_thio_) and thiocarbonyl-containing compounds (thio_) etc etc. I tried to give the root name to the group that may be most causing the problem…hence “cyano_imine_”.
Let’s look at a few more examples:
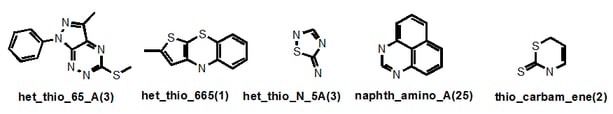
“Het_thio_” describes any heterocycle where a suspected problematic (endo- or exo-) sulfur is present and the ring fusion is described also. But if an exocyclic alkene was present for example, this would take priority. Where an S-N bond is present, this significantly problematic feature gains its own nomenclature (“thio_N”) as in “het_thio_N_5A”. Sometimes I have decided to give readily recognized systems that are present in several different classes their own name, as in “naphth”. There are seven naphthalene-containing classes in our filters, four of which belong to the napth_amino_ grouping. So in this case, you wouldn’t find these under the “het_” root name. Similarly with “thio_carbam_ene” – the thiocarbonyl takes priority.
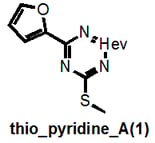
I do not claim absolute rigour and I note that thio_pyridine_A below should really be a “het_thio_6_furan”. But you get my drift.
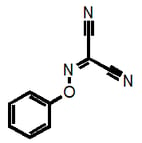
Why am I going into this detail? Because there is a danger that these filters could unintentionally be used for good rather than evil, that’s why. For example, a given researcher may have this as a hit:
This would not be recognized by our filters as they are currently strictly defined and could therefore lead to the conclusion that “we have shown that this compound is NOT a PAIN”. But it clearly belongs to the highly problematic cyano_imine_ class of compounds and itself would plausibly be a problematic PAIN. If the academic researcher is familiar with my naming system, they can readily find this out (parenthetically I point out that neither may some real nasties be recognized if the functional group filter that we also provide isn’t used prior).
So why didn’t I broaden the definitions: for example, to remove all sulfur-containing compounds and all cyanoimines? In the alpha version of our filters I went down this line in many cases, but I found that I started to dislike the look of almost everything and increasingly large numbers of possibly benign systems were being removed (one wonders if this natural human tendency might sometimes extend to big pharma and result in diminished access to genuine screening diversity). For example, I was considering defining all hydrazones as problematic because they appeared to associate frequently with problematic compounds and they are also very common compounds. However, on further inspection, it was clear that certain sub-classes of hydrazones were causing the major problems. Indeed, one of our most successful medicinal chemistry programs started with a hydrazone-containing hit which we would never have found because an alternative linkage between the different binding elements was not commercially represented. And we were able to fairly quickly optimize out the PK-liable hydrazone linkage.
So in the end, I made an effort to adopt a purely evidence-based approach and define structural classes as well as possible to give more specific sub-classes with significantly higher EBAELL percentages.
Finally, a technicality. The 480-odd classes we define are not necessarily all PAINS. It is a significant indictment for a compound class to be defined as a PAIN and so at this stage I have required that not only must a class be problematic in our assays, but there must be evidence in the literature that similar compounds are hitting different targets using different assays (an intriguing situation where privileged structures and PAINS may intermingle. And in some ways also a paradox, because by these criteria I therefore require false positives to be published in the first place!). I undertook a literature search in my paper only for the most common and obvious likely candidates and so have elevated only a handful of problematic classes to PAIN status and I name these in the main body of our paper. For this reason the numerous other classes we list in the supplementary figure I am currently calling putative PAINS.
I hope that our filters may help a little the academic drug discovery researcher and indirectly lead towards enrichment in the literature of better points for efficient therapeutic development.
For now, it is regrettably the case that the scientific literature is being flooded with misinformed publications.
But those who live in glass houses should not throw stones. Some of these PAINS have fooled me and my colleagues in the recent past. They could fool anyone. Well, almost anyone. Many would not fool big pharma (though we've a reason to believe a few of our lesser-known PAINS may be new to some of them).
If you’ve successfully licensed your biologically active compounds to big pharma (not necessarily a smaller biotech and certainly not a spin-out, depending on who is on their SAB with big pharma experience), you can be at least be fairly sure that you’re onto something special. This is a rare and significant event for the translational researcher and in utter contrast to the potential worthlessness of an unlicensed provisional patent. In fact, the vast numbers of patents falsely based on PAINS as revealed by a quick SciFinder search is somewhat disheartening and represents an enormous cost to those Institutions.
We argue at our Institute – with some success - that this is more than the equivalent of a biologist’s Nature paper. Conversely, if you can’t license your IP, perhaps you should be asking for honest answers why not.
Licensed IP should be true succour for your promotion committee. More importantly, perhaps this is the ultimate tangible metric for measuring your translational track record.
This blog is authored by members of the CDD Vault community. CDD Vault is a hosted drug discovery informatics platform that securely manages both private and external biological and chemical data. It provides core functionality including chemical registration, structure activity relationship, chemical inventory, and electronic lab notebook capabilities!
CDD Vault: Drug Discovery Informatics your whole project team will embrace!
Other posts you might be interested in
View All Posts
CDD Vault Updates
4 min
July 12, 2016
GSK's Kinetoplastid Research Data Set Available FREE Within CDD Vault
Read More
CDD Blog
2 min
October 14, 2014
Get Ready for CDD’s First Live Webinar Event!
Read More
CDD Blog
2 min
March 20, 2014
CDD’s 10th Anniversary Community Meeting Agenda Now Online
Read More