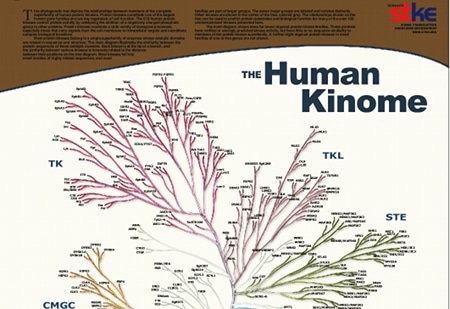
used with permission from http://kinase.com/human/kinome/
Many CDD users are familiar with ChEMBL, a monumental database of bioactive drug-like small molecules that contains medicinal chemistry, bioinformatics and bioassay data integrated from a wide variety of sources (the literature, deposited data sets, other bioassay databases). The database is maintained by the European Bioinformatics Institute (EBI), based on an award from The Wellcome Trust, resulting in the creation of the ChEMBL chemogenomics group at EBI, led by John Overington.
Among the many valuable resources available at ChEMBL is the Kinase SARfari subset. As stated at the ChEMBL website, the Kinase SARfari is “an integrated chemogenomics workbench focused on kinases. The system incorporates and links kinase sequence, structure, compounds and screening data”. A highly useful resource, this database makes available a large amount of SAR data for the kinase active compounds against a wide range of kinases in a broad array of assays, much of it manually mined and curated from the literature.
While this database and associated interface are extremely useful to kinase biochemists, it is not as conducive to SAR assessment as a kinase medicinal chemist might like. To make this data available in the CDD interface, we took the core table of Kinase SARfari and merged it with another key table from the ChEMBL database, providing a field describing the assay utilized in each record in much greater detail than is available in native SARfari. We have now made this merged dataset available via the CDD interface.
In this new dataset, we have created 400 protocols, where each protocol represents one of the 400 kinases in the SARfari database. This organization allows users to search by molecule, substructure, kinase target, or assay result. This allows one to investigate SAR of a variety of molecules against a single target, or cross-reactivity of a series of compounds against multiple kinases.
In addition to this kinase bioactivity data, SARfari makes available two other significant datasets. The first is known as the “Starlite ADMET” dataset. These data are molecule specific, but not target related. Instead, they generally represent in vitro or in vivo PK or tox data on kinase inhibitors. These data have been extracted into a single CDD protocol called ADMET Data. The second dataset is known as the “Starlite Functional” data. These data are again molecule (not target) specific, and generally represent the results of a wide variety of in vivo or ex vivo experiments. These data have been extracted into the CDD protocol Functional Data.
This blog is authored by members of the CDD Vault community. CDD Vault is a hosted drug discovery informatics platform that securely manages both private and external biological and chemical data. It provides core functionality including chemical registration, structure activity relationship, chemical inventory, and electronic lab notebook capabilities!
CDD Vault: Drug Discovery Informatics your whole project team will embrace!
Other posts you might be interested in
View All Posts
CDD Blog
8 min
October 22, 2018
Why Is Publicly Available Data Important And How Is It Being Used?
Read More
CDD Blog
3 min
June 17, 2015
CDD Awarded $1.5 Million Grant to Simplify Encoding of Bioassays with Automated Annotation Technology
Read More
CDD Blog
3 min
July 8, 2013
GSK Published Kinase Inhibitor Set (PKIS) Now Available in the CDD Vault
Read More


