July 19, 2019
CDD Awarded Phase 1 SBIR Grant on an Intelligent Chemical Structure Browser for Drug Discovery and Optimization

CDD Awarded Phase 1 SBIR Grant on an Intelligent Chemical Structure Browser for Drug Discovery and Optimization
Burlingame, California - July 15, 2019 - Collaborative Drug Discovery, provider of CDD Vault web-based drug discovery informatics platform, announced they won a competitive, peer-reviewed Phase 1 SBIR grant from NIH NCATS, titled: "Intelligent Chemical Structure Browser for Drug Discovery and Optimization".
Collaborative Drug Discovery, Inc. (CDD) proposes to develop a novel intelligent data browser that will enable medicinal chemists developing new drug compounds to more efficiently browse and organize experimental data in an intuitive way that - for the first time - matches how they mentally map the relationships among the closely similar molecules that comprise a chemical lead series.
A key innovation of our approach is a novel lattice-based method that we have conceived to represent the relationship among molecules in a lead series. Naive algorithms are far too slow, requiring hours to construct a proper relationship diagram among a few hundred molecules. We estimate that our approach will scale to thousands, and ultimately millions, of compounds in just a few minutes (on the same hardware), enabling us to create a delightful user experience that will present intuitive, automated chemical hyperlinking while hiding the internal complexity. The lattice representation will also enable new types of sophisticated, automated structure-activity relationship (SAR) analyses of lead series data, such as generalized forms of Free-Wilson analysis and automated suggestions of which missing experimental data would best strengthen the understanding of the SAR. In Phase 1, we will develop the lattice representation and associated algorithms and demonstrate that they scalably support all the operations needed to realize the vision of the software. In Phase 2, we will build the complete application, develop further analysis methods, and test it with external partners.
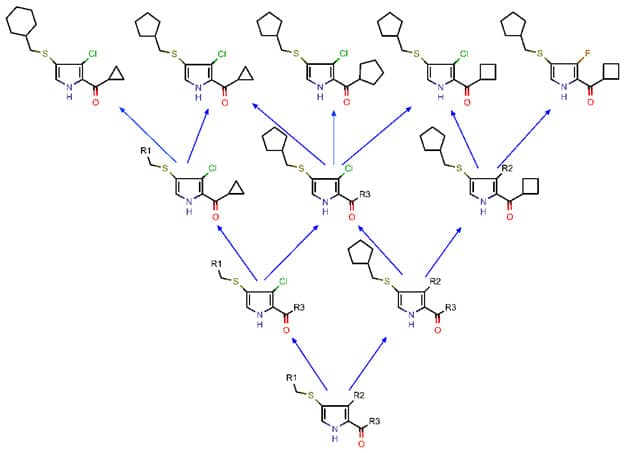
Figure 1: Example of a few hypothetical molecules in a lead series (top row) and a small portion of the internal semilattice data structure (not exposed to users in the UI) that represents their relationship to each other and to other molecules that aren't shown. Arrows denote the substructure-superstructure relationship. The semilattice will be generated by a recursive procedure.
Navigating through and extending a lead series to discover the optimal drug candidate to advance into animal studies and clinical trials comprises a critical stage of the drug discovery pipeline: the success of large subsequent investments depends on making the right decision. This stage also especially emphasizes creative and intuitive thinking. Existing software that assists scientists engaged in this task tabulates data in formats that make it difficult to assemble and compare the essential data needed to rapidly explore ideas about how to further optimize promising candidates. Our proposed intelligent browser represents a highly innovative solution that will essentially "hyperlink" chemical space. It will allow chemists to navigate easily among compounds following the same pathways that lead from one compound to the next in the mental models that they intuitively map in their heads, and to instantly create compact tables containing just the right experimental data to answer the questions that emerge from exploring any particular pathway. These capabilities in turn will enable medicinal chemists to make better decisions about which new compounds to synthesize and submit for expensive assays that experimentally assess their pharmacological properties. Since it is feasible to test only a limited number of compounds, better scientific decisions at this stage can significantly increase the chances that a drug candidate will successfully emerge through the clinical pipeline as an approved drug, and lead to a drug that is more effective and more safe.
Specific Aims for Phase 1 are to:
- Develop an algorithm to efficiently construct the lattice that organizes a set of input molecules according to structure. We also aim to demonstrate fast performance on sets of up to 10,000 molecules and that the resulting lattice obeys essential mathematical properties of a partially ordered set, so that the algorithm constructs the same diagram for any set of molecules, independent of the order in which they are processed.
- Develop a second key algorithm that can automatically identify SAR neighborhoods in the lattice.
- Define the user interface concept and show that the data representation supports all the planned functions.
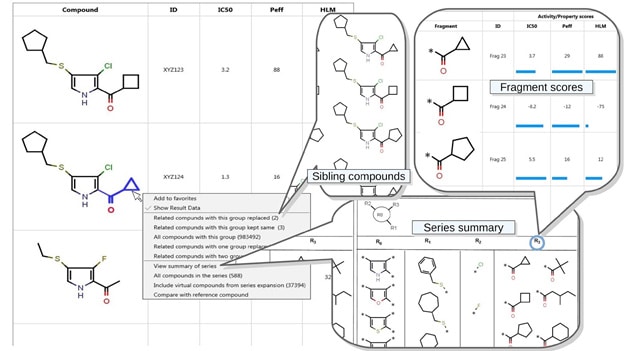
Composite of several screens from a preliminary UI Mock-up
- Build the complete application and test its performance with one or more external partners.
- Further optimize the key algorithms to support millions of compounds with reasonable computing resources.
- Develop advanced capabilities, such as automated, generalized Free-Wilson regression.
ABOUT THIS GRANT
The Small Business Innovation Research (SBIR) is part of a program to enable sharing of biological data. Award Number #1R43TR002699-01 from National Center for Advancing Translational Sciences as described on NIH Reporter supports this project. This content is solely the responsibility of the authors and does not necessarily represent the official views of the National Center for Advancing Translational Sciences or the National Institutes of Health.
ABOUT COLLABORATIVE DRUG DISCOVERY, INC.
CDD's (collaborativedrug.com) flagship product, "CDD Vault®", is used to manage chemical registration, structure-activity relationships (SAR), and securely scale collaborations. CDD Vault® is a hosted database solution for secure management and sharing of biological and chemical data. It lets you intuitively organize chemical structures and biological study data, and collaborate with internal or external partners through an easy to use web interface. Available modules within CDD Vault include Activity & Registration, Visualization, Inventory, and ELN.
A complete list of more than 60 publications and patents from CDD can be found online on our resources page at https://www.collaborativedrug.com/publications-and-resources/.
Other posts you might be interested in
View All Posts
CDD Blog
3 min
October 23, 2020
Collaborative Drug Discovery Announces a Grant Supplement Building on a Decade Long Commitment to Global Tuberculosis Drug Discovery Scientists
Read More
CDD Blog
4 min
April 8, 2021
CDD Awarded $1.5M Phase 2 SBIR Grant on an Intelligent Chemical Structure Browser for Drug Discovery and Optimization
Read More
CDD Blog
4 min
December 5, 2022
Collaborative Drug Discovery Announces New NIH Grant to Develop Cloud Workspaces for Drug Discovery
Read More


