In this guest blog, Dr. Andrew Norris describes how the UCLA – Center for Medical Countermeasures against Radiation (CMCR) implemented CDD.
The chief aim of the UCLA CMCR is the discovery and development of novel small molecule drug leads that serve as radiation mitigators in addition to advancing our understanding of radiation toxicity and lethality in humans.
CDD has been very important for our program overall and has been implemented in the UCLA Radiation Protection Screening Core as part of that effort. Within this core (Figure 1), phenotypic and target based screen results are entered into the CDD database linking a biological result to a chemical structure. The data is mined for trends such as structure activity relationships, substructure analysis, and other determining information that may be inside or outside the UCLA program. Thoroughly tested chemical entities (leads) are then synthesized and this is then further explored via proteomics, chemical informatics and analogue synthesis as a platform to support the management of the screening information as well as the discovery of mechanism of action. Multiple lead candidates have been tested in animals. This data is all compiled back into the database. Over the years, the breadth of data associated with the compounds has become significant and several successful lead molecules have been produced.
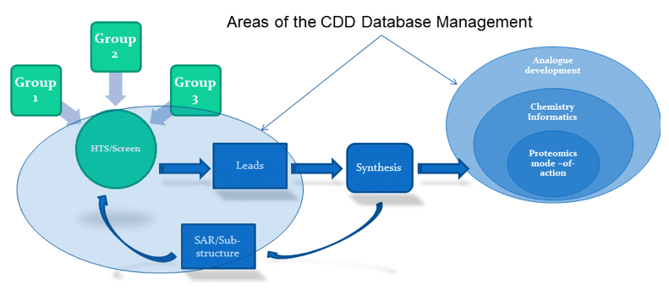
Figure 1: Schematic of the overall strategic approach of the UCLA-CMCR screening program. CDD plays an integral part in the storage, management, processing and social networking of groups involved in the effort.
What we have found particularly useful about the CDD platform is that we were able to network the multiple research groups from entirely different locations and disciplines with the small molecules being the common language between them all. Another useful aspect of the software is that the database serves to distinguish what aspects of the small molecules analyzed are most effective or responsible for mitigating cellular damage/death from ionizing radiation. This has been a critical aspect of our SAR studies and also important for ruling out false negatives in our data.
Furthermore, this platform allows for access to novel structures containing desired characteristics trough commercial and private sources, the database is flexible to include almost any type of biological data input associated with a small molecule being examined. To date, over 15 research groups from different backgrounds are networked through the database and over 100,000 compounds have been investigated for activity.
This blog is authored by members of the CDD Vault community. CDD Vault is a hosted drug discovery informatics platform that securely manages both private and external biological and chemical data. It provides core functionality including chemical registration, structure activity relationship, chemical inventory, and electronic lab notebook capabilities!
CDD Vault: Drug Discovery Informatics your whole project team will embrace!
Tag(s):
CDD Blog


