We are very excited to bring you this release - it has been brewing for some time and contains many new additions to CDD Vault. We have completed a major upgrade to the infrastructure that runs CDD Vault, with a focus on improving performance, availability, and disaster recovery capabilities. In addition, CDD Vault now includes a handful of highly sought after features, based on all the feedback we collected from you, our users. Which one of these have you been anticipating?
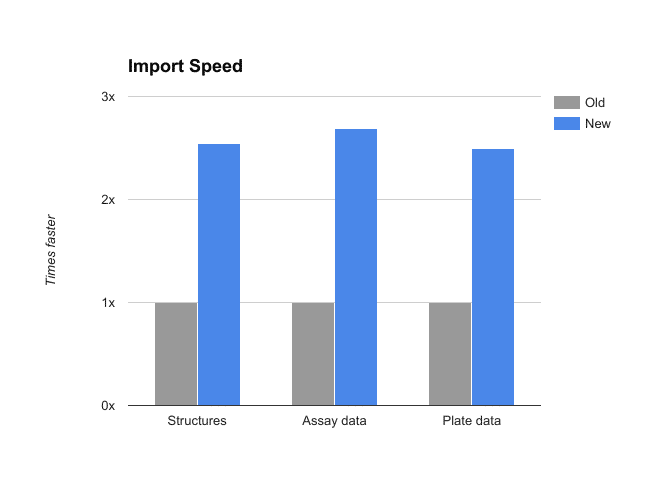
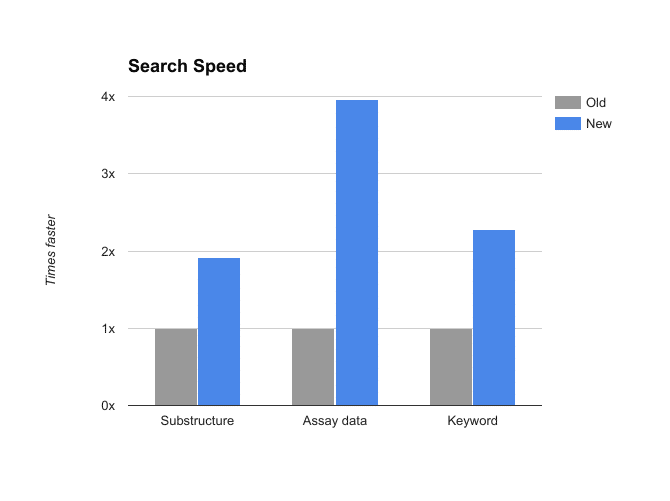 CDD Vault should be approximately twice as fast as it was last week, whether you are registering molecules, adding plate and assay data, or searching results.
CDD Vault should be approximately twice as fast as it was last week, whether you are registering molecules, adding plate and assay data, or searching results.
 For our existing customers - we have created a migration tool that will allow you to convert existing protocol readouts to a pick list format. Please contact
support@collaborativedrug.com to find out how. We are working on bringing the same functionality to batch fields - stay tuned!
Learn more about pick lists.
For our existing customers - we have created a migration tool that will allow you to convert existing protocol readouts to a pick list format. Please contact
support@collaborativedrug.com to find out how. We are working on bringing the same functionality to batch fields - stay tuned!
Learn more about pick lists.
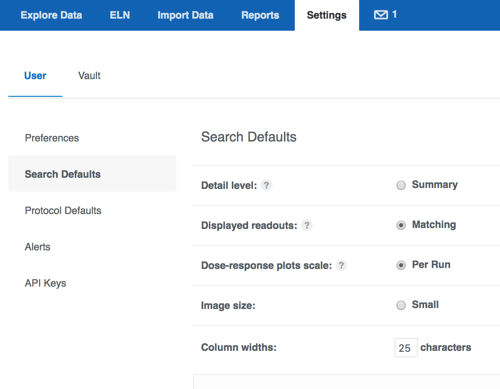 These search defaults include molecule and batch related fields, as well as structure and chemical properties. Many users have asked us to extend this ability to create custom defaults for protocol data as well. We are very pleased to introduce this option in this release!
These search defaults include molecule and batch related fields, as well as structure and chemical properties. Many users have asked us to extend this ability to create custom defaults for protocol data as well. We are very pleased to introduce this option in this release!
 Learn more about search and protocol defaults.
Learn more about search and protocol defaults.
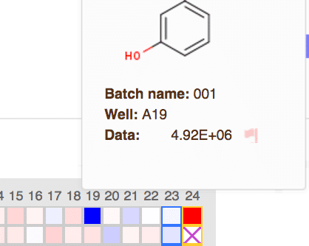
 Alternately, you may now mark outliers directly from heat-maps! You can visually identify outliers easily on a heat-map, and now you do not need to navigate away from this view to remove them from calculations. Just click on the well you wish to mark and click the red flag to remove the data point. Use the readout definition drop-down to mark additional outliers. Notice how the marked well is crossed out - these values will have special styling on the search results table as well, and will not be used in any calculations.
Alternately, you may now mark outliers directly from heat-maps! You can visually identify outliers easily on a heat-map, and now you do not need to navigate away from this view to remove them from calculations. Just click on the well you wish to mark and click the red flag to remove the data point. Use the readout definition drop-down to mark additional outliers. Notice how the marked well is crossed out - these values will have special styling on the search results table as well, and will not be used in any calculations.
 Thank you for reading to the end. We hope to deliver the most user-friendly and advanced data management tool to you, and look forward to hearing your feedback anytime to
support@collaborativedrug.com.
Thank you for reading to the end. We hope to deliver the most user-friendly and advanced data management tool to you, and look forward to hearing your feedback anytime to
support@collaborativedrug.com.
- Performance of our upgraded infrastructure is roughly 2 to 4 times faster on imports and searches!
- Pick lists (controlled data types for protocol readouts) provides controlled assay vocabulary.
- User-based search defaults let individuals customize their search results display.
- User-based protocol defaults let individuals customize their default protocol readout definitions.
- Outlier marking is now available directly from heat-maps and across all results of a search.

 CDD Vault should be approximately twice as fast as it was last week, whether you are registering molecules, adding plate and assay data, or searching results.
CDD Vault should be approximately twice as fast as it was last week, whether you are registering molecules, adding plate and assay data, or searching results.
Pick list (controlled data type for protocol readouts)
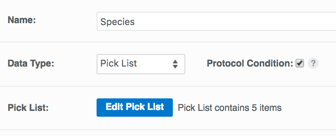
The ability to control the values that are permitted in your protocols has always been important. After all the "garbage in - garbage out" rule applies to CDD Vault just like any other database. Thus far, this control was exercised at the import file template level, where users were responsible for ensuring the correct spelling of species, tissues, phenotypes, and any other assay conditions. But as diligent as our users are, it is better to have the application perform data validation automatically based on the permissible set of values. The new pick list data type allows vault administrators and protocol owners to do just that: define data dictionaries and control the vocabulary of permissible values for individual protocol readouts. Especially useful when defining protocol conditions, the new data type can be selected when creating a new readout. Once selected, there is an option to add pick list items either one at a time or by pasting a list. You can edit the pick list to add or remove items, and you can even "archive" an item, preventing new entries but maintaining the option to search on previously entered data. For our existing customers - we have created a migration tool that will allow you to convert existing protocol readouts to a pick list format. Please contact
support@collaborativedrug.com to find out how. We are working on bringing the same functionality to batch fields - stay tuned!
Learn more about pick lists.
For our existing customers - we have created a migration tool that will allow you to convert existing protocol readouts to a pick list format. Please contact
support@collaborativedrug.com to find out how. We are working on bringing the same functionality to batch fields - stay tuned!
Learn more about pick lists.
User-based search defaults
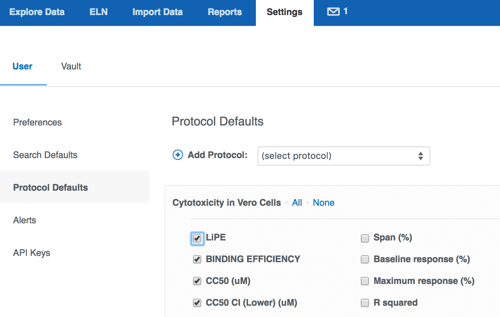
Everyone is used to clicking "customize your report" to adjust their search results table. In the past, vault administrators could configure certain fields to be displayed automatically when a new search was run. We have now moved this option to individual users - now everyone may define for themselves the exact set of fields they'd like to display by default on their new searches. The new option resides on the Settings page, User settings, and should look familiar - we use the same "customize your report" interface. These search defaults include molecule and batch related fields, as well as structure and chemical properties. Many users have asked us to extend this ability to create custom defaults for protocol data as well. We are very pleased to introduce this option in this release!
These search defaults include molecule and batch related fields, as well as structure and chemical properties. Many users have asked us to extend this ability to create custom defaults for protocol data as well. We are very pleased to introduce this option in this release!
User-based protocol defaults
Similar to molecule and batch search defaults, users are now able to configure their individual default display options for protocols. Once set, the protocol defaults will be automatically applied when a new protocol search is run, or when the protocol is added to the results table via the "Customize your report" panel. The new option resides on the Settings, User settings, and should once again look familiar. Learn more about search and protocol defaults.
Learn more about search and protocol defaults.
Outlier marking
Marking outliers has become more streamlined. On the search page the link now resides directly in the table header, allowing you to mark outliers across all search result rows and columns. If your link is greyed out, make sure to include raw or normalized data in the table. Thank you for reading to the end. We hope to deliver the most user-friendly and advanced data management tool to you, and look forward to hearing your feedback anytime to
support@collaborativedrug.com.
Thank you for reading to the end. We hope to deliver the most user-friendly and advanced data management tool to you, and look forward to hearing your feedback anytime to
support@collaborativedrug.com.Other posts you might be interested in
View All Posts
CDD Vault Snack
2 min
June 16, 2019
CDD Vault Snack #2 - Search and Protocol Defaults
Read More
CDD Vault Updates
2 min
November 22, 2013
CDD Vault Update: Search page updates
Read More
CDD Vault Updates
2 min
December 27, 2022
CDD Vault Update (December 2022 [#2]): New Protocol Search & Structure Editor Features
Read More


