For 3D Vision, Mine PDB Ligand Structures in CDD Vault
Guest blog from CDD Advocate: Dr. Dale Cameron
Part 2 of 2
In a recent blog post, CDD announced the addition of the latest ligand structures from the Protein Data Bank (PDB) into the CDD Vault®. The expanded availability of this unique and useful data is indeed exciting news, but just what can the scientific community actually DO with this data?
Figure 1 illustrates one of many examples of what can now be done from within CDD Vault. Substructure, exact and similarity searches are easily performed within CDD Vault. To investigate an interesting backbone, simply perform a query to find related structures in the PDB and one click brings you all the ligand information including the PDB link to structure complexes that are available. As an example, let's look at trifluoroazepine (TFP), a typical antipsychotic of the phenothiazine chemical class.
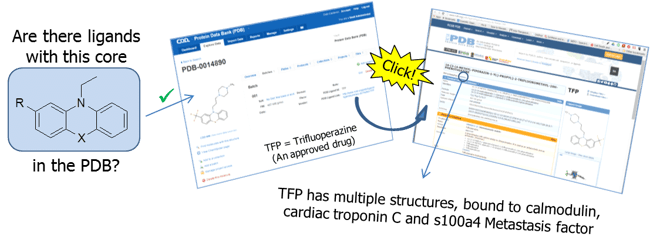

Figure 1: Example of Ligand data stored in CDD Vault with an excerpt of associated data from the PDB
You are not limited to just searching structures. Text searches for targets, PDB codes, or ligands, as well as filters for structure types (x-ray, NMR, electron microscopy) and resolution or deposit dates are among the many options that have been curated for your convenience. For example, a simple search for structure titles including “HIV Integrase” reveals 67 unique ligand records. In this one simple query, a set of ligands that bind to HIV Integrase can be put into a CDD Vault "collection" and shared with your colleagues through your own private CDD Vault.
And, since all this new information is shared as a public dataset, any search of a CDD Vault of interest (public or private), combined with a search of the PDB vault, will now provide structural information about the ligand, without you even expressly knowing the structure existed. So cool!
For example, if you were interested in identifying new trypanosome inhibitors, a search of public data sets in CDD Vault reveals over 300K compounds have been tested in an HTS screen against T. cruzi.1 Combining these screening results with a search of the PDB compounds with at least one crystal structure reveals 527 unique compounds of interest (without bias towards their respective target) all from a single "AND" search within CDD Vault. Refining that search by searching for hits with “Trypanosoma cruzi” in the PDB structure title provides three structural leads: fluconazole (complex with sterol 14-alpha-demethylase (CYP51)), N-acetyl-2,3-dehydro-2-deoxy-neuramic acid (complex with trans-sialidase) and 5-fluoro-orotic acid (complex with dihydroorotate dehydrogenase). Figure 2 shows a snapshot of the data available for fluconazole. It is now easy to find not only interesting starting points for design of new inhibitors but also a way to identify known and interesting protein targets. Storing these three compounds as a collection in your CDD Vault, allows you to share your results with your colleagues and perform further searches to uncover more structural information in the PDB.
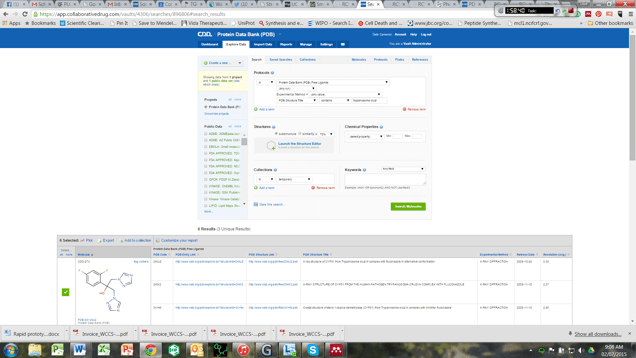
Figure 2: Example data for fluconazole found thru queries within CDD Vault: PDB code, experimental method, original release date and resolution combined with links to the PDB entry page or PDB structural download.
A subsequent search of the PDB returns all of the structures associated with those three leads, to give a sense of how this information can be further pursued. There are other potentially useful data in the PDB, following up on the initial observation from within CDD Vault following the convenient links. For example, fluconazole has also been crystallized with Trypanosome bruceii, Mycobacterium tuberculosis and Leishmania infantum CYP51. A comparison of all these structures may reveal common design elements to begin novel inhibitor design targeting a broad set of human pathogenic organisms.
Similarly, as final examples (and you’ll find your own examples based on your scientific interests), N-acetyl-2,3-dehydro-2-deoxy-neuramic acid has been crystallized with many other targets including other trypanosome sialidases, viral and bacterial neuraminidases as well as human sialidase. 5-fluoro-orotic acid has also been crystallized with E. coli and human dihydroorotase domains. This cross-species information and comparison to human binding details is potentially useful in designing specific and potentially less toxic, novel compounds.
From each PDB CDD Vault entry, a link to the original PDB page as well as a direct download link (for the actual PDB structure file) is available. This provides direct access to more detailed information and 3D structure files, both of which are instantly available. From the PDB, much more information is available, including 3D viewing capabilities, citations, detailed experimental, protein sequence, etc.
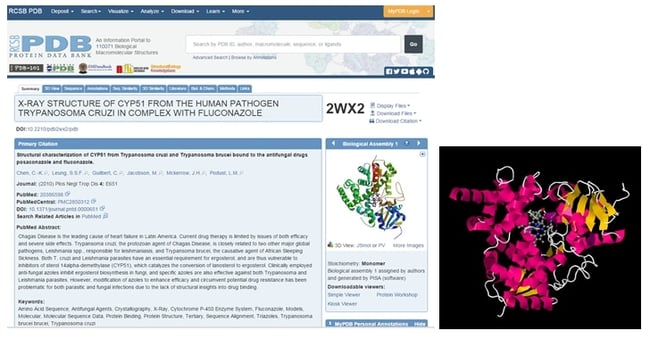
Figure 3: PDB page for fluconazole bound to T. cruzi (2WX2) and 3D view of the structure. (Source1, Source2)
As mentioned in our initial blog announcement, the PDB is maintained by the Research Collaboratory for Structural Bioinformatics (RCSB) consortium and contains a wealth of interesting information, connected to many other data sources. With the publicly available PDB CDD Vault dataset, we have coupled the inherent power of collaborative data analysis in CDD Vault with the immensely valuable PDB datasets all at your fingertips. As always, it is available to the community read-only freely in CDD Public...and with privacy, import, export, plus advanced analytics and unique collaborative capabilities commercially in CDD Vault. If you have suggestions for other key data of broad scientific interest that could be stored CDD Vault, please let us know.
1. Ekins, S.; Lage de Siqueira-Neto, J.; McCall, L.-I.; Sarker, M.; Yadav, M.; Ponder, E. L.; Kallel, E. A.; Kellar, D.; Chen, S.; Arkin, M.; Bunin, B. A.; McKerrow, J. H.; Talcott, C. PLoS Negl. Trop. Dis. 2015, 9, e0003878.
This blog is authored by members of the CDD Vault community. CDD Vault is a hosted drug discovery informatics platform that securely manages both private and external biological and chemical data. It provides core functionality including chemical registration, structure activity relationship, chemical inventory, and electronic lab notebook capabilities!
CDD Vault: Drug Discovery Informatics your whole project team will embrace!
Other posts you might be interested in
View All Posts
CDD Vault Updates
3 min
August 26, 2015
Want 3D Vision? Protein Data Bank (PDB) Ligand Structures are now LIVE in CDD Vault® (Part 1 of 2)
Read More
CDD Blog
1 min
June 26, 2012
Virtual Pharma, Externalization, FIPNet, VIPCO – like gravity, you can fight it, but you’ll lose (Part 2 - Simplicity)
Read More
News
2 min
November 3, 2014
CDD Announces Partnership With Ryoka Systems
Read More


