The big story in this release is the new CDD Portal, which allows you to search across all of your CDD Vaults and available data sets at once. Prior to this release, if you had data in multiple vaults, you could only search within each vault individually. This release also includes a much requested smaller enhancement to saved searches. If you specify display options for your search and then save it, the display options are saved along with the search, so that if you run the saved search again you get the columns you wanted in the order specified. Very soon (within a week) we will also retain your chosen display options when you refine a search. This is the second release in a four-part series of "Projects" releases, which overall will provide the ability to work on live, shared, selected data with selected collaborators, the logical next step from sharing readonly snapshots with selected collaborators (which is current functionality). We won't achieve this vision until the fourth release, but each release is a step in a logical progression. We'll take a break from this progression for the next release, resuming in April, to allow time for user feedback. The next release will focus on raw data processing (calculation of percent inhibition, percent of control, B-scores, etc.). This release includes an important terminology change. Within the product we have renamed "Group" to "Vault" to be consistent with our marketing literature. We're still getting used to it, too. In general we think this term is an improvement because the term, "group", can be ambiguous.
 We are currently extending this to retain display options when a search is refined (e.g. another protocol criterion is added). Also, we plan to restrict private data sets shared from saved searches to publish only the readout definitions and user-defined fields selected in the display options. Both of these small enhancements will be released next week.
We are currently extending this to retain display options when a search is refined (e.g. another protocol criterion is added). Also, we plan to restrict private data sets shared from saved searches to publish only the readout definitions and user-defined fields selected in the display options. Both of these small enhancements will be released next week.
 Users who only belong to one non-sandbox vault will be taken to their vault dashboard, but they can still navigate up to the portal dashboard by clicking the link at the top of the screen, "Up to Your CDD Dashboard".
Users who only belong to one non-sandbox vault will be taken to their vault dashboard, but they can still navigate up to the portal dashboard by clicking the link at the top of the screen, "Up to Your CDD Dashboard". 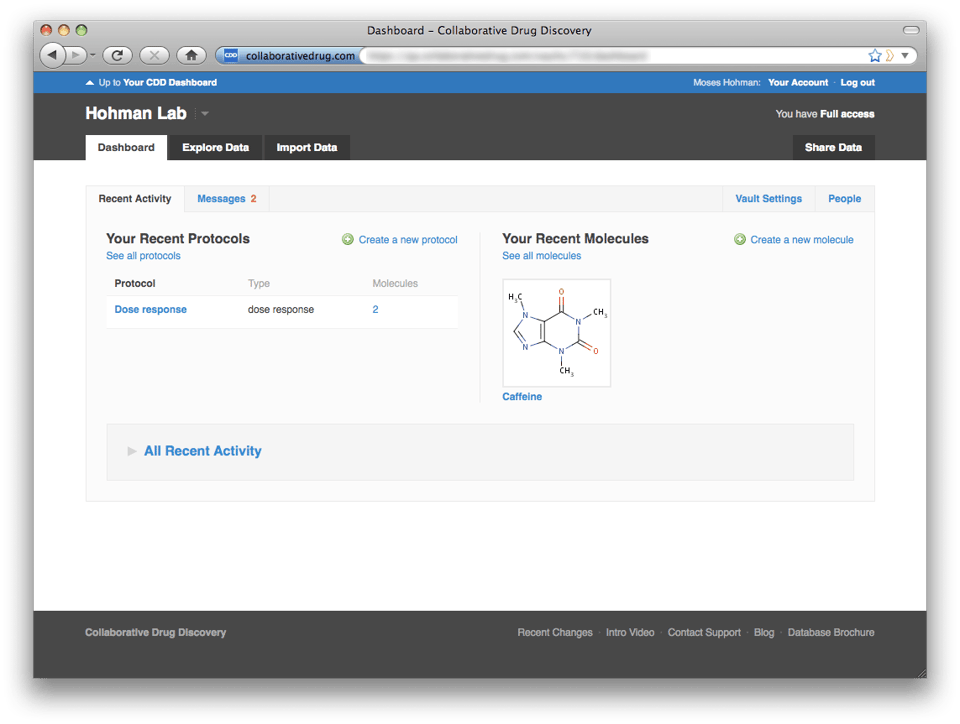 The portal level dashboard gives you a convenient overview of activity in all of your vaults. By default each vault is collapsed to hide details and preserve screen real estate. However, if you expand a vault section, the system will remember that you prefer to have that vault expanded.
The portal level dashboard gives you a convenient overview of activity in all of your vaults. By default each vault is collapsed to hide details and preserve screen real estate. However, if you expand a vault section, the system will remember that you prefer to have that vault expanded.
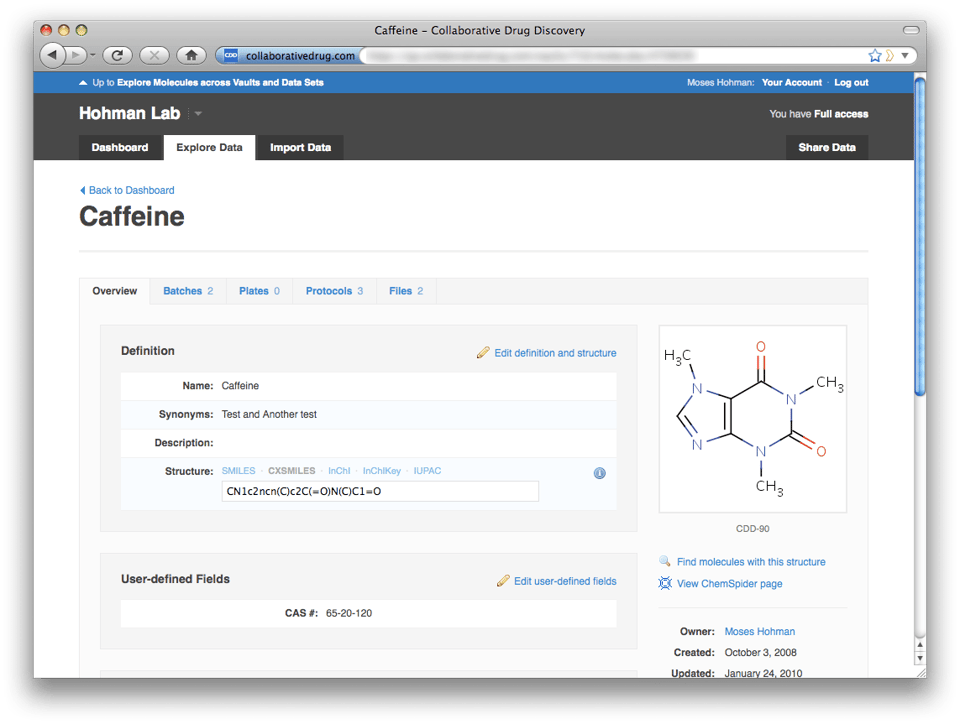 Descending back into a vault from a particular molecule or protocol page is also easy. Just click on the blue vault name ("Hohman Lab" below) on the upper right. Note that the portal top-level navigation tabs consist only of "Dashboard" and "Explore Data"—"Import Data" and "Share Data" are missing. These functions exist only within a vault, where it's very clear where you're importing data to and whose data you're sharing.
Descending back into a vault from a particular molecule or protocol page is also easy. Just click on the blue vault name ("Hohman Lab" below) on the upper right. Note that the portal top-level navigation tabs consist only of "Dashboard" and "Explore Data"—"Import Data" and "Share Data" are missing. These functions exist only within a vault, where it's very clear where you're importing data to and whose data you're sharing. 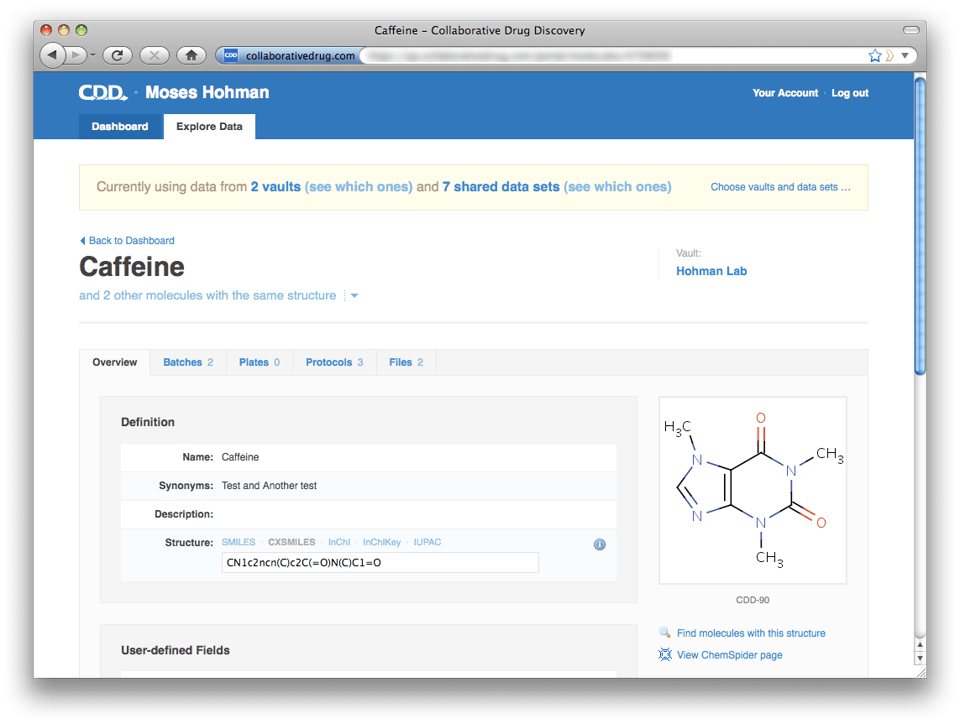
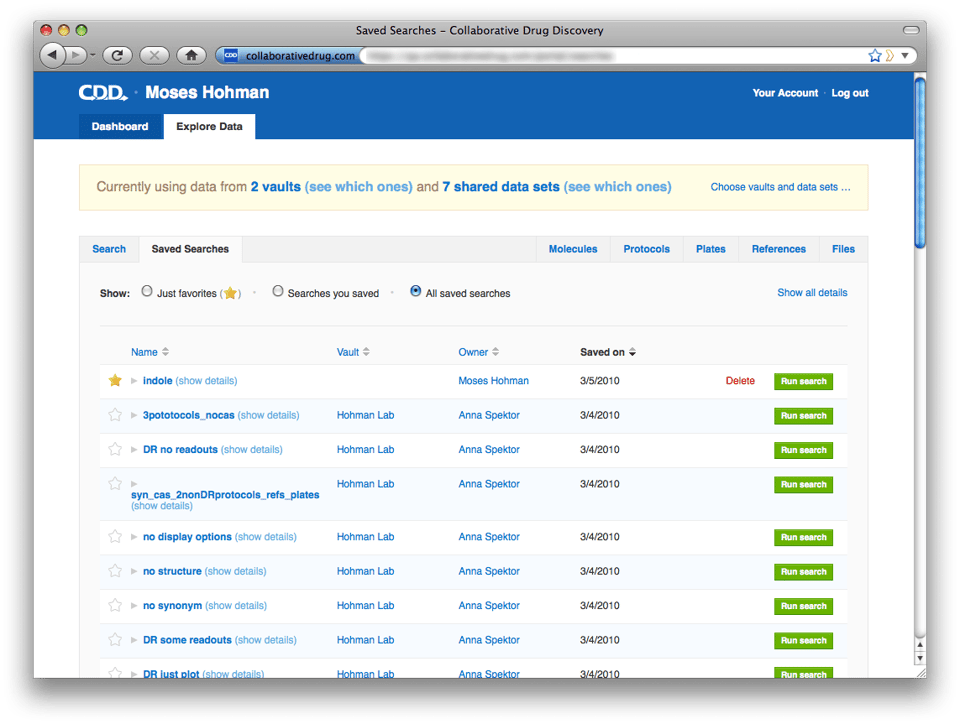
Saved Display Options
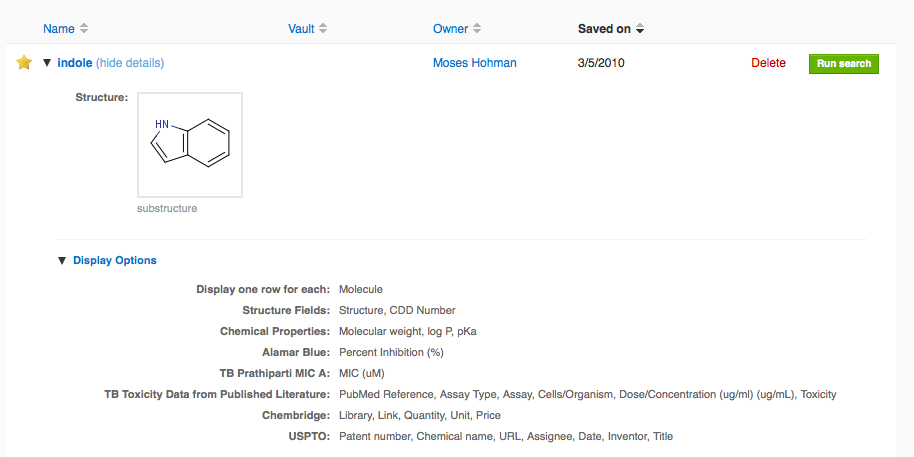
As mentioned above, if you specify display options on a search and then save it, the display options are saved along with your search. If you run this saved search, your column selections and ordering will be preserved. To see the display options saved with a search, just expand the search in the newly redesigned saved searches page and then expand the display options: We are currently extending this to retain display options when a search is refined (e.g. another protocol criterion is added). Also, we plan to restrict private data sets shared from saved searches to publish only the readout definitions and user-defined fields selected in the display options. Both of these small enhancements will be released next week.
We are currently extending this to retain display options when a search is refined (e.g. another protocol criterion is added). Also, we plan to restrict private data sets shared from saved searches to publish only the readout definitions and user-defined fields selected in the display options. Both of these small enhancements will be released next week.
CDD Portal
Users with multiple CDD Vaults will most appreciate the new CDD Portal functionality, because it allows them to search across all of their vaults simultaneously. We have kept users with a single vault in mind, however.Dashboard
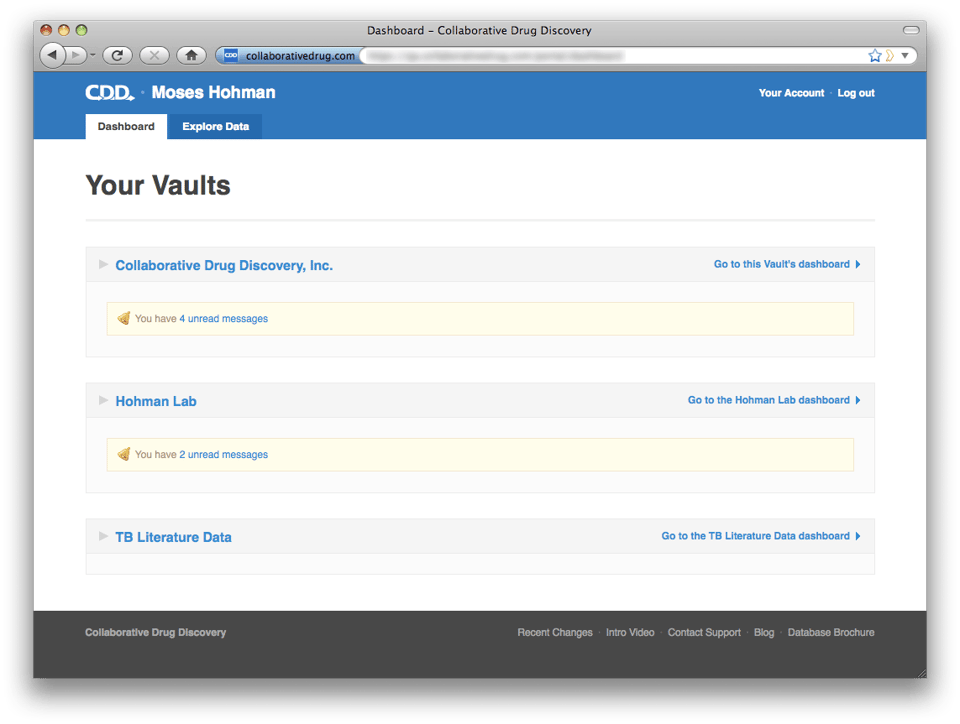
If you belong to more than one non-sandbox CDD Vault, when you log in to CDD you will be taken to your portal-level dashboard instead of a vault dashboard. Users who only belong to one non-sandbox vault will be taken to their vault dashboard, but they can still navigate up to the portal dashboard by clicking the link at the top of the screen, "Up to Your CDD Dashboard".
Users who only belong to one non-sandbox vault will be taken to their vault dashboard, but they can still navigate up to the portal dashboard by clicking the link at the top of the screen, "Up to Your CDD Dashboard".  The portal level dashboard gives you a convenient overview of activity in all of your vaults. By default each vault is collapsed to hide details and preserve screen real estate. However, if you expand a vault section, the system will remember that you prefer to have that vault expanded.
The portal level dashboard gives you a convenient overview of activity in all of your vaults. By default each vault is collapsed to hide details and preserve screen real estate. However, if you expand a vault section, the system will remember that you prefer to have that vault expanded.
Navigation
One major change with this release is that you can no longer search public and privately shared data sets at the vault level. All cross-vault and cross-data set mining is done at the portal level. Similarly, importing data can only be done at the vault level. We have introduced this separation to make it very clear how to add data to a particular vault and how to check what data is contained in that vault. To make it easy to pop up to the portal context should you find yourself in a vault wanting to search across data sets, on every page at the vault level there is a white link at the top of the page in the blue header bar that will take you to the corresponding page at the portal level. Descending back into a vault from a particular molecule or protocol page is also easy. Just click on the blue vault name ("Hohman Lab" below) on the upper right. Note that the portal top-level navigation tabs consist only of "Dashboard" and "Explore Data"—"Import Data" and "Share Data" are missing. These functions exist only within a vault, where it's very clear where you're importing data to and whose data you're sharing.
Descending back into a vault from a particular molecule or protocol page is also easy. Just click on the blue vault name ("Hohman Lab" below) on the upper right. Note that the portal top-level navigation tabs consist only of "Dashboard" and "Explore Data"—"Import Data" and "Share Data" are missing. These functions exist only within a vault, where it's very clear where you're importing data to and whose data you're sharing. 
Search and Vault/Data Set Selection
Now for the most exciting part. With the ability to search across all your vaults also comes a revamped vault and data set selection page and saved searches page. The Search page looks very similar. One important difference: When specifying protocol search criteria, you now have the option to specify, for example, "any Hohman Lab protocol". You can use criteria like this to find molecules that have been screened in common across your vaults or across data sets. To choose vaults and data sets to search across, click "Choose vaults and data sets..." in the yellow banner. We changed the design of this page considerably, we hope you find it more intuitive than the previous design. The redesigned Saved Searches page shows all searches saved across all your vaults, plus searches you've saved at the portal level. To help you organize this list, we've provided the ability to mark useful searches as favorites. You can filter the list of searches to include only your favorites, only searches saved by you, or view the entire list. Each search listing is brief by default, because you will probably recognize by name the searches important to you. If you need to find a search saved by someone else, and you don't know its name, you can click "Show all details" on the upper right and browse through the expanded list.
Miscellaneous
The most serious bug fixed concerned searches that specified both a structure (for similarity or substructure search) and another criterion, such as a keyword. If the structure search by itself returned more than the maximum number of hits (more than 5,000), it was possible for the query to return fewer than the actual number of results for the combined criteria (structure plus keyword). Thanks to Erin Bradley for noticing this and pointing it out to us! This release includes two important performance improvements. Data import is now almost two times as fast (88% faster in our tests). We have also fixed a major performance problem with some multi-protocol searches, which used to time out but now run in seconds. We fixed a number of small issues with data import related to errors or inconsistencies in the file causing the entire file to become "unparseable". These problems are now reported as errors on the QC Report. We also now report an imported file's inconsistent batch vendor values as errors, rather than just use the last value mentioned in the file. For those uploading and importing large files, we have added a small but not yet ideal feature to speed up the upload process. You can now upload a CSV or SD file compressed with the gzip algorithm, and it will function just like a normal file. Gzipped SD files are typically 10% of their uncompressed size, so this speeds up the upload by a factor of 10 (because it's limited only by the bandwidth between your computer and CDD). Apple and Linux users have built-in access to the gzip program for compression. Windows users can download the open source software utility, 7-Zip, to compress data files with the gzip algorithm. We plan to extend this functionality in the future to include .zip files as well (.zip files are more complicated because they can contain multiple files and subdirectories).Other posts you might be interested in
View All Posts
CDD Vault Updates
2 min
June 17, 2025
CDD Vault Update (June #2 2025): Bulk Register Sequences with Custom Monomers, Set X-axis Ranges in Plots, Filter AI Bioisostere Suggestions, Inventory Debits from ELN
Read More
CDD Blog
1 min
June 17, 2025
CDD Vault Curves vs. GraphPad Prism: Which Tool is Best for Early-Stage Drug Discovery Data Analysis?
Read More
CDD Blog
1 min
June 17, 2025
Why We Committed to Amazon Web Services — And What It Means for You
Read More


