We introduce CDD Vision, a new analysis and visualization product, predictive model building within CDD Vault, and a half-dozen smaller enhancements. We are excited to see what you can achieve with the tools in this release.
 This release allows you to use chemical properties and readouts within a single protocol in your calculations. Upcoming releases will allow you to combine readout definitions from separate assays in one calculation.
This release allows you to use chemical properties and readouts within a single protocol in your calculations. Upcoming releases will allow you to combine readout definitions from separate assays in one calculation.  The interactive formula builder makes it easy to define your calculations, and it should feel familiar to Excel users. The best part is that, once created, the calculated results are updated automatically when new data are imported into a protocol, or chemical properties change because a structure is edited.
The interactive formula builder makes it easy to define your calculations, and it should feel familiar to Excel users. The best part is that, once created, the calculated results are updated automatically when new data are imported into a protocol, or chemical properties change because a structure is edited.
 The model will automatically score all compounds in the project you selected when creating it. You can subsequently share it with other projects to score other molecules. Each molecule receives a relative score, applicability metric, and maximum similarity number.
The model will automatically score all compounds in the project you selected when creating it. You can subsequently share it with other projects to score other molecules. Each molecule receives a relative score, applicability metric, and maximum similarity number.  It is worth pointing out that the maximum similarity value is independent of the Bayesian model, and provides a way for you to perform a similarity search using all active compounds at once. The models feature is very much in beta, and will be evolving significantly in the coming year. We'd love to hear what you want to be able to do with models, to help guide that evolution.
It is worth pointing out that the maximum similarity value is independent of the Bayesian model, and provides a way for you to perform a similarity search using all active compounds at once. The models feature is very much in beta, and will be evolving significantly in the coming year. We'd love to hear what you want to be able to do with models, to help guide that evolution.



Calculations within a protocol – part of CDD Vision
This is the introductory release of CDD Vision, an optional extension to CDD Vault. When CDD Vision is enabled in your vault, you can perform custom calculations on protocol readouts, chemical properties, and numeric constants. Later releases will bring a dynamic visualization suite for decision making support. To start using CDD Vision right away, contact info@collaborativedrug.com for the early adopter program. Calculations are defined as a new type of readout definition within a protocol. The interactive formula builder makes it easy to define your calculations, and it should feel familiar to Excel users. The best part is that, once created, the calculated results are updated automatically when new data are imported into a protocol, or chemical properties change because a structure is edited.
The interactive formula builder makes it easy to define your calculations, and it should feel familiar to Excel users. The best part is that, once created, the calculated results are updated automatically when new data are imported into a protocol, or chemical properties change because a structure is edited.
Model builder – part of CDD Vision
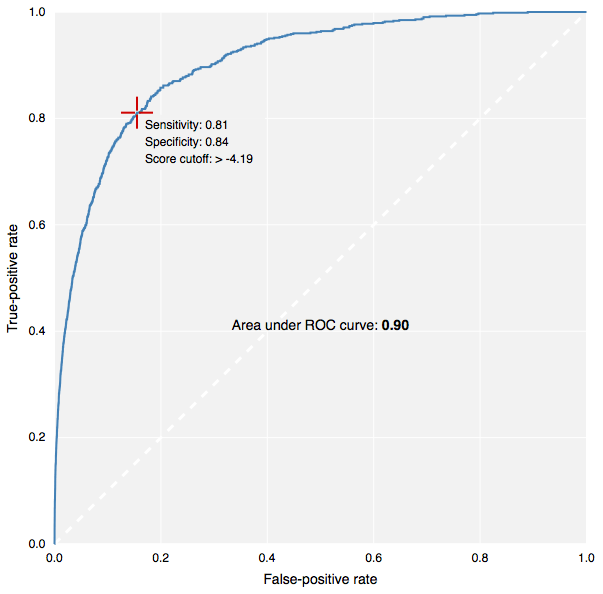
This is also the introductory release of CDD Models, which is included in your CDD Vision subscription. Please contact info@collaborativedrug.com to learn how to start using it. Creating a model only requires that you choose the assay to use as the training set and select the active compounds. A Bayesian statistical model is built using the FCFP6 fingerprints of your structures, and is then used to rank compounds that you have not screened yet. The model is stored as a type of protocol, and provides a ROC plot so you can gauge its effectiveness. The model will automatically score all compounds in the project you selected when creating it. You can subsequently share it with other projects to score other molecules. Each molecule receives a relative score, applicability metric, and maximum similarity number.
The model will automatically score all compounds in the project you selected when creating it. You can subsequently share it with other projects to score other molecules. Each molecule receives a relative score, applicability metric, and maximum similarity number.  It is worth pointing out that the maximum similarity value is independent of the Bayesian model, and provides a way for you to perform a similarity search using all active compounds at once. The models feature is very much in beta, and will be evolving significantly in the coming year. We'd love to hear what you want to be able to do with models, to help guide that evolution.
It is worth pointing out that the maximum similarity value is independent of the Bayesian model, and provides a way for you to perform a similarity search using all active compounds at once. The models feature is very much in beta, and will be evolving significantly in the coming year. We'd love to hear what you want to be able to do with models, to help guide that evolution.
Other notable changes
Export plate maps
When you prepare cherry-picked or dilution plates, you can easily download parent compound plate maps directly from the “Plates” tab. Just search for the plate name(s), then click to export plate maps. Exported files contains the plate and well position, with molecule and batch assignment.
Multiple file attachments
For messages, molecules, plates, runs, references and protocols, multiple files can be uploaded in a single step by selecting more than one file with your computer’s file manager application (Explorer on windows, Finder on Mac). Images of all attachments are also rendered as previews once uploaded.Printer friendly plots
Dose-response plots may be printed as monochrome (gray-scale) images by selecting the “monochrome” option when exporting to Excel.
Updated dose-response calculation
Activity points above the intersection are no longer required for dose-response calculation. For example, an IC50 can still be reported when no responses exceed 50%.Updated API GET call
GET Molecule now returns IUPAC and InChI structures, in addition to other molecule properties and structure formats.pKa type
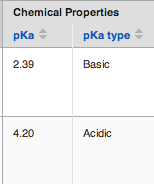
CDD reports the strongest acidic or basic pKa value. We have added a field to indicate whether the calculated pKa value is for an acidic or basic center.


