SARvision|SM and CDD Vault
SARvision|SM has functionality to connect with CDD Vault and import data stored in Vault for easy Structure Activity Analysis (or Sequence Structure Analysis for SARvision|Biologics). SARvision|SM allows you to identify chemotypes automatically, group your molecules around scaffolds in a hierarchical tree, create easily R-group tables, grid views, perform match molecular pairs analysis, enumerate libraries, visualize data with plots that relate to chemotypes, filter tables by selected property ranges, search libraries with multiple queries at once, seamless connectivity with Word and Excel for your reports. AMEDEO, an add-on to SARvision|SM, provides tools for lead optimization using machine learning.A few simple steps show how to get access to your CDD Vault account and then pull data relevant to your analysis into SARvision|SM
1) Log in to the CDD Vault and go to Settings. Under API Keys | Add a new Key to use with SARvision. Copy this key so that it can be pasted into SARvision in the next step. This key contains all the information to allow SARvision|SM to have access to you CDD Vault database. Note that it can be deleted to remove access at a later date.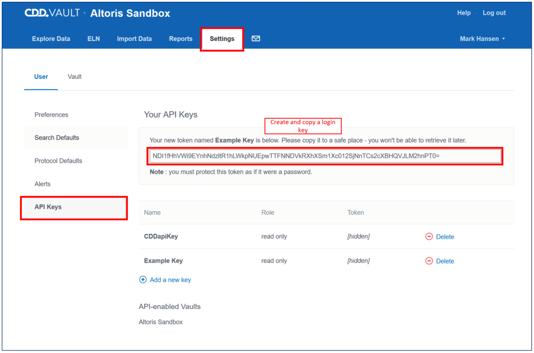 2) In SARvision|SM, under File->Import from->CDD Vault… paste the API token that you copied in step one into the correct box as shown. The Update button will reload the available saved queries in CDD Vault that you can access. If you do not have any, log onto CDD Vault and create some. In the ChemApps CDD Vault connector, select one of the queries and press the Import button.
2) In SARvision|SM, under File->Import from->CDD Vault… paste the API token that you copied in step one into the correct box as shown. The Update button will reload the available saved queries in CDD Vault that you can access. If you do not have any, log onto CDD Vault and create some. In the ChemApps CDD Vault connector, select one of the queries and press the Import button. 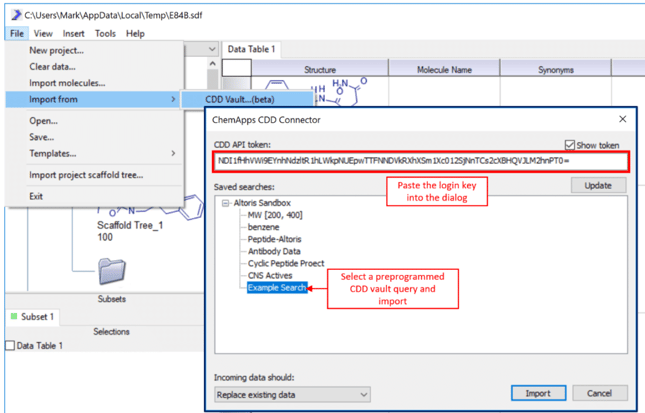 3) Once you have loaded some data, right click on the Data Table tab and use the Columns… control to clean up the table and show only the relevant columns.
3) Once you have loaded some data, right click on the Data Table tab and use the Columns… control to clean up the table and show only the relevant columns. 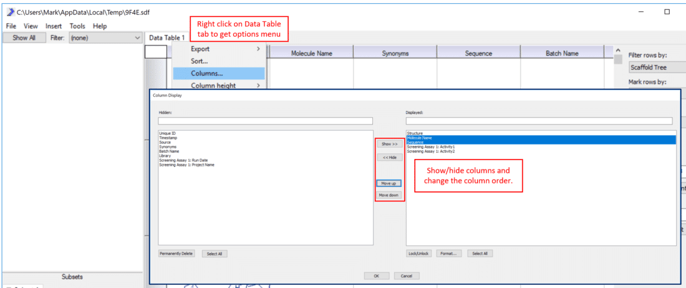 4) Right click on the Scaffold Tree workspace (upper left) to get a tree menu. Select Identify scaffolds… and use the default settings to generate relevant chemical cores or scaffolds that represent this particular dataset.
4) Right click on the Scaffold Tree workspace (upper left) to get a tree menu. Select Identify scaffolds… and use the default settings to generate relevant chemical cores or scaffolds that represent this particular dataset. 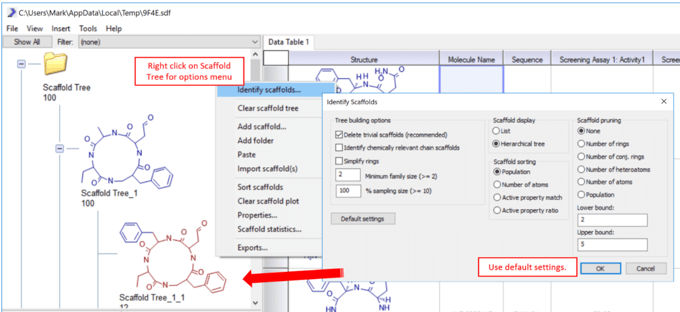 5) The user can edit any scaffold in the tree by right clicking on that scaffold and selecting Edit scaffold… The user can also drag and drop scaffolds, delete scaffold and manually draw in scaffold using this menu.
5) The user can edit any scaffold in the tree by right clicking on that scaffold and selecting Edit scaffold… The user can also drag and drop scaffolds, delete scaffold and manually draw in scaffold using this menu. 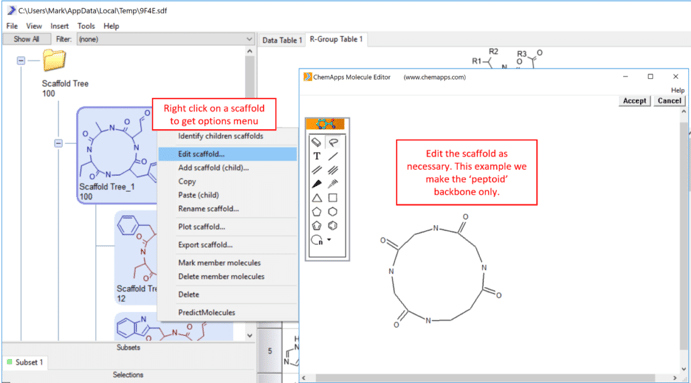 6) Additional view of the data can be added under main menu->Insert. In this example an R-Group table is added. To trigger rebuilding of the tables, double click on any scaffold. This will load the molecules that belong to that scaffold, reorient them relative to the scaffold and color code the molecules based on the scaffold.
6) Additional view of the data can be added under main menu->Insert. In this example an R-Group table is added. To trigger rebuilding of the tables, double click on any scaffold. This will load the molecules that belong to that scaffold, reorient them relative to the scaffold and color code the molecules based on the scaffold. 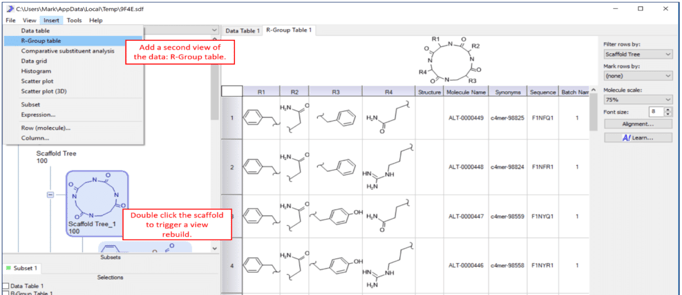 Now you have access to a very large number of tools for lead optimization, library design or other tasks common in small molecule drug discovery.
Now you have access to a very large number of tools for lead optimization, library design or other tasks common in small molecule drug discovery.
This opens a number of options for you to analyze data...
Grid Views: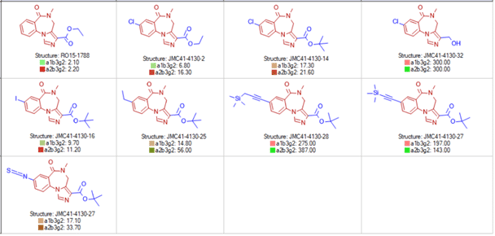 RxR Tables:
RxR Tables: 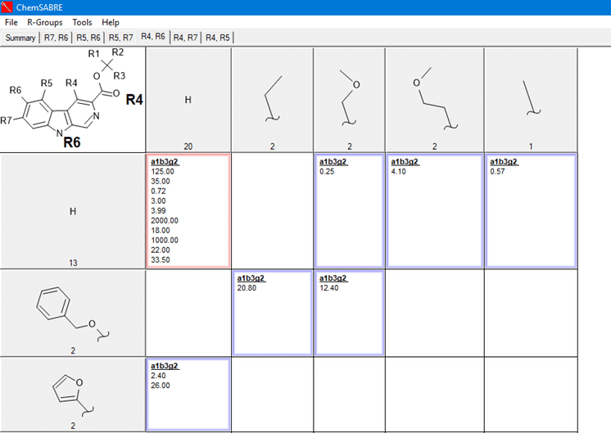 Matched Molecular Pairs:
Matched Molecular Pairs: 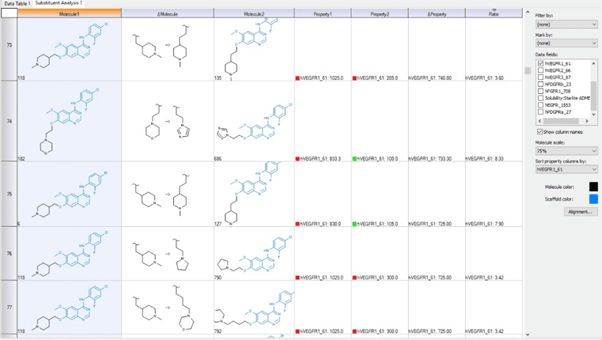 Several graphing options including Scaffold plots:
Several graphing options including Scaffold plots: 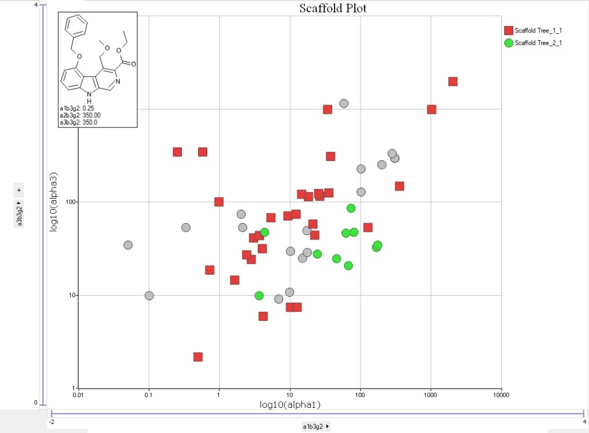 And add the power of Machine Learning for lead optimization of activity or ADME related properties using AMEDEO.
And add the power of Machine Learning for lead optimization of activity or ADME related properties using AMEDEO. 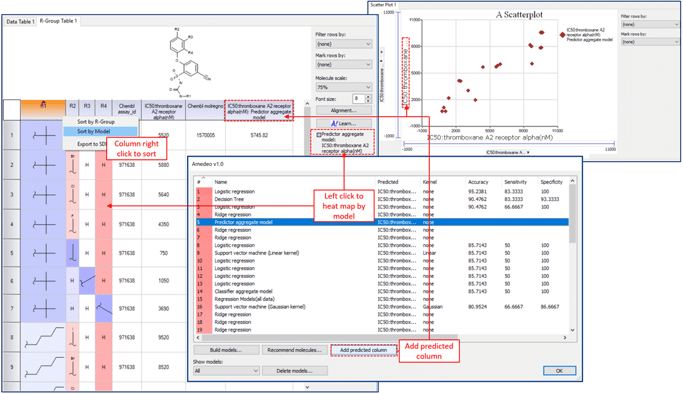 CDD Vault also connects seamlessly with SARvision|Biologics as well, the program provides a platform to read-in and organize data on biological polymers.The program can be viewed as a smart spreadsheet that simplifies work with biopolymers including peptides, proteins, nucleic acids, chemically modified residues and unnatural amino acids. SARvision|Biologics provides a number of tools to relate sequence to activity. Altoris, the makers of SARvision |SM have posted a short video on You-Tube that shows the steps for integration beetween CDD Vault and SARvision |SM. As always, feel free to contact CDD Support for assistance.
CDD Vault also connects seamlessly with SARvision|Biologics as well, the program provides a platform to read-in and organize data on biological polymers.The program can be viewed as a smart spreadsheet that simplifies work with biopolymers including peptides, proteins, nucleic acids, chemically modified residues and unnatural amino acids. SARvision|Biologics provides a number of tools to relate sequence to activity. Altoris, the makers of SARvision |SM have posted a short video on You-Tube that shows the steps for integration beetween CDD Vault and SARvision |SM. As always, feel free to contact CDD Support for assistance.Other posts you might be interested in
View All Posts
CDD Vault Snack
3 min
July 20, 2021
Vault Snack #14 – Sharing Strategies for Data You’ve Imported into CDD Vault
Read More
CDD Vault Snack
2 min
December 13, 2021
Vault Snack #15 – Importing Data into CDD Vault Temporarily Locks your Vault
Read More
CDD Vault Updates
3 min
July 25, 2021
CDD Vault Update (July 2021 [#2]): Solvent Workflow and New ELN API Endpoints
Read More


