 Barry Bunin, PhD
Barry Bunin, PhD
Founder & CEO
Collaborative Drug Discovery
Drug Discovery Industry Roundup with Barry Bunin
"AlphaFold accelerates artificial intelligence powered drug discovery: efficient discovery of a novel CDK20 small molecule inhibitor" is the title of a recent study published in the Royal Society of Chemistry journal Chemical Science. Researchers at the University of Toronto, Stanford University, and Insilico Medicine developed a potential treatment for hepatocellular carcinoma (HCC) with an AI drug discovery platform called Pharma.AI. Any potential drug would, of course, need to go through clinical trials before use. "This paper is further evidence of the capacity for AI to transform the drug discovery process with enhanced speed, efficiency, and accuracy," Michael Levitt, a Nobel Prize winner in chemistry, said. Co-author, Alan Aspuru-Guzik, Professor of Chemistry and Computer Science at the University of Toronto forecasted: "If one uses a generative model targeting an AI-derived protein, one can substantially expand the range of diseases that we can target. If one adds self-driving labs to the mix, we will be in uncharted territory. Stay tuned!"
* * *
"Meta AI Unlocks Hundreds of Millions of Proteins to Aid Drug Discovery." The Wall Street Journal carries that headline about Facebook parent company Meta Platforms Inc.'s new tool that predicts the structure of hundreds of millions of proteins using artificial intelligence. A tool, called ESMFold is fed the amino acids sequences for protein's with known structures. The AI model then learns how to fill in the sections in the sequence that were blank or hidden. Once it had generated a complete sequence, ESMFold then learns the relationship between known protein sequences and structures that are already well-understood by scientists to predict the structures of new ones. Meta scientists say ESMFold's strength is the speed at which it can predict protein structures, allowing researchers to search through large genetic databases to find possible applications in medicine, health, food and the environment. While the article quotes researchers who prefer the greater accuracy they see with AlphaFold, others see the two programs working well together, with each tool having strengths that can lead to new discoveries. Andrew Ferguson, co-founder of Chicago-based biotech Evozyne and Associate Professor of Molecular Engineering at the University of Chicago, said: "They are complementary," adding that the Meta AI model "was a really elegant idea."
Researchers say it promises to deepen scientists' understanding of biology, and perhaps speed the discovery of new drugs. Since a picture is worth 1000 words, here is an image of the ESM Metagenomic Atlas covering both knowing and unknown protein structures and their nearest neighbor:
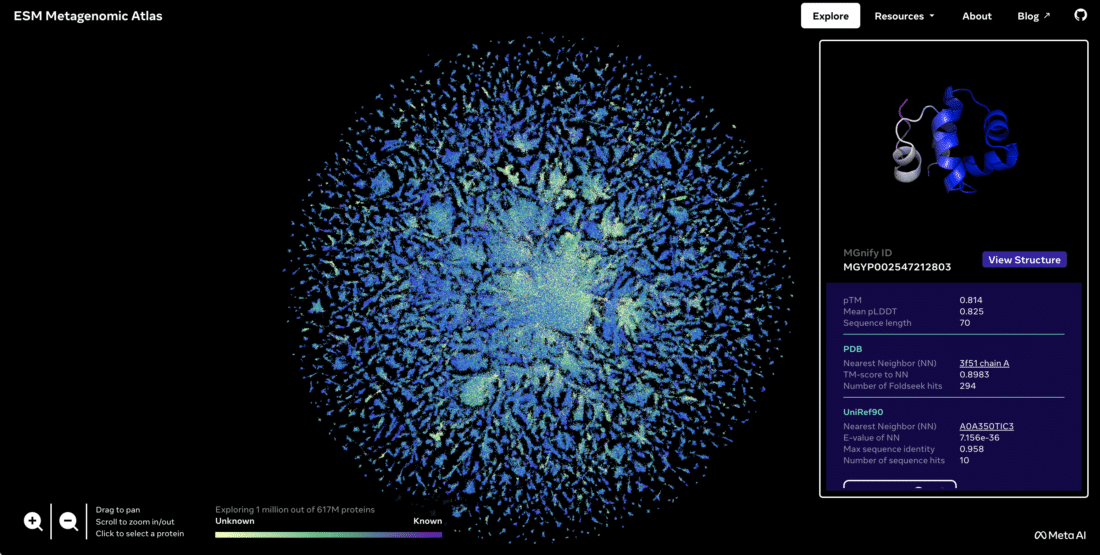
In addition to the excitement and buzz around Facebook's response to Google's DeepMind AlphaFold splash on their blog…those interested in reading the science under the hood can check out these free preprints on bioRxiv "Language Models of Protein Sequences at the Scale of Evolution Enable Accurate Structure Predictions" and "Evolutionary-scale Prediction of Atomic Level Protein Structure with a Language Model." These recent papers are timely given the current excitement (and hype) around ChatGPT4 and other LLMs (Large Language Models).
* * *
Harvard Law on "How Artificial Intelligence is Revolutionizing Drug Discovery." Now even Harvard Law is showing an interest in AI and drug discovery. A recent article provides an overview of milestones in AI-enabled drug discovery, and the value of using an AI-first approach. Declaring "The future of AI-enabled drug discovery is already here," the author writes: "If current trends continue, it will only be a matter of time before the drugs we take are no longer designed by people, but by machines. With the promise of lower costs and shorter development timelines, AI-enabled drug discovery holds massive potential to increase the accessibility of drugs and to treat presently incurable conditions. However, it also opens the floodgates to a host of unresolved issues relating to, e.g., intellectual property rights, the risk of technological misuse, and the continued assurance of drug safety and efficacy in this new era."
* * *
Everything Everywhere All at Once. Too bad that phrase was taken by the Oscar-winning movie. Otherwise, Derek Lowe might have used it as a subtitle for his recent Science blog titled "The Wild World of RNA-Binding Proteins." He discusses what he describes as "some neat chemical biology that has revealed some unexpected behavior in cancer cells," pointing to a collaboration between groups at UCSD, Scripps-La Jolla, and Janssen searching for small molecules that affected the expression of the androgen receptor which is implicated in several tumor varieties. After describing surprises the researchers found, he summarizes: "To me, this all illustrates the benefits of coming at these pathways from every direction you can think of. There are more mechanisms that can operate in the living cell than we can anticipate at the whiteboard, and the phenotypic approach of "try it and see what happens" is much better suited, frankly, to our level of understanding of tumor cell biology." He concludes his piece: "Well, there are more things in heaven, hell, and the cell than are dreamt of in our philosophy, and that's why you shouldn't talk yourself out of interesting experiments. We don't know enough to go around ruling a lot of things out. That may be a bit of a blow to the ol' ego, but it's the truth. Abandon your pride - we have a lot to learn!"
=
Barry A. Bunin, PhD, is the Founder & CEO of Collaborative Drug Discovery, which provides a modern approach to drug discovery research informatics trusted globally by thousands of leading researchers. The CDD Vault is a hosted biological and chemical database that securely manages your private and external data.
Other posts you might be interested in
View All Posts
CDD Vault Updates
2 min
June 17, 2025
CDD Vault Update (June #2 2025): Bulk Register Sequences with Custom Monomers, Set X-axis Ranges in Plots, Filter AI Bioisostere Suggestions, Inventory Debits from ELN
Read More
CDD Blog
1 min
June 17, 2025
CDD Vault Curves vs. GraphPad Prism: Which Tool is Best for Early-Stage Drug Discovery Data Analysis?
Read More
CDD Blog
1 min
June 17, 2025
Why We Committed to Amazon Web Services — And What It Means for You
Read More


